Glycans are fundamental to life processes such as energy provision, cellular signaling, and immune recognition. These diverse functions derive directly from their unique three-dimensional structures. However, research into glycan structure and function has significantly lagged behind that of nucleic acids and proteins. The primary challenges lie in the diversity of monosaccharide molecules, each featuring multiple chiral centers, the complexity and variability of glycosidic linkages in glycan formation, and their inherent structural flexibility. These factors collectively render high-resolution three-dimensional structural determination exceptionally difficult, severely limiting our understanding of their structures and functions.
"CryoSeek" strategy proposed by Nieng Yan's team employs cryogenic electron microscopy as a discovery tool for unknown biological macromolecules without prior knowledge. This strategy may therefore represent "forward structural biology", a structure-first paradigm for biological discovery. In their initial study, the team combined cryogenic electron microscopy with AI-facilitated autobuilding tools and bioinformatic analyses to characterize a glycofibril, TLP-1, in water samples from Tsinghua lotus pond, and speculated its origin and potential function. Subsequent work further identified a novel glycofibril structure, TLP-4. Its core comprises a linear chain of tetrapeptide repeats, coated by a dense layer of glycans. Each tetrapeptide repeat consists of a conserved 3,4-dihydroxyproline (DiHyp) with 3-OH and 4-OH being highly O-glycosylated, as well as an adjacent conserved O-glycosylated serine or threonine. These findings not only reveal the crucial role of glycans in the structural assembly of biological macromolecules but also provide novel insights for discovering and elucidating natural glycan structures [1, 2, 3].
However, a fundamental challenge arises when determining structures using cryogenic electron microscopy: the imaging process loses "absolute hand" information. Consequently, three-dimensional reconstruction yields a density map assigned randomly to one of the two possible "mirroring configurations". For proteins, this ambiguity can be resolved by using prior knowledge, such as the fact that α-helices are exclusively right-handed. However, glycofibrils are primarily composed of glycans that lack α-helices. Moreover, both D-sugars and L-sugars coexist in nature. Consequently, even at atomic resolution, their absolute hand cannot be directly determined. This ambiguity is analogous to the loss of "north-south orientation on map", making it exceedingly difficult to build a correct atomic model.
The conventional approach to this problem was the "tilt-pair" imaging method, which, however, requires prior knowledge of a reference structure and the acquisition of two images of the same area. For highly heterogeneous samples containing hundreds of distinct glycofibrils, such as those collected directly from natural freshwaters, this method is practically inapplicable.
On October 1, 2025 (Beijing Standard Time), the teams led by Nieng Yan and Mingxu Hu, together with their collaborators, posted two preprints on BioRxiv. They are entitled "CryoSeek identification of glycofibrils with diverse compositions and structural assemblies" and "Absolute hand determination of glycofibrils from natural sources in cryo-EM", respectively.

Link to full text: https://doi.org/10.1101/2025.09.30.679562

Link to full text: https://doi.org/10.1101/2025.09.30.679555
Continuing the "Glycans from the Lotus Pond" initiative, the team subsequently identified five additional glycofibrils through further data processing. This work underscores the remarkable diversity of glycofibrils in natural environments and highlights the crucial role of glycans in structural assembly. Concurrently, to tackle the persistent challenge of determining absolute hand in glycofibrils, the team developed a new method named Ahaha. This approach requires only mono-tilt cryogenic electron microscopy to determine the absolute hand of natural glycofibrils efficiently and accurately. The method is now available as an online service (Fig. 1).
Fig. 1 Ahaha online service page (https://cryoseek.org/ahaha)
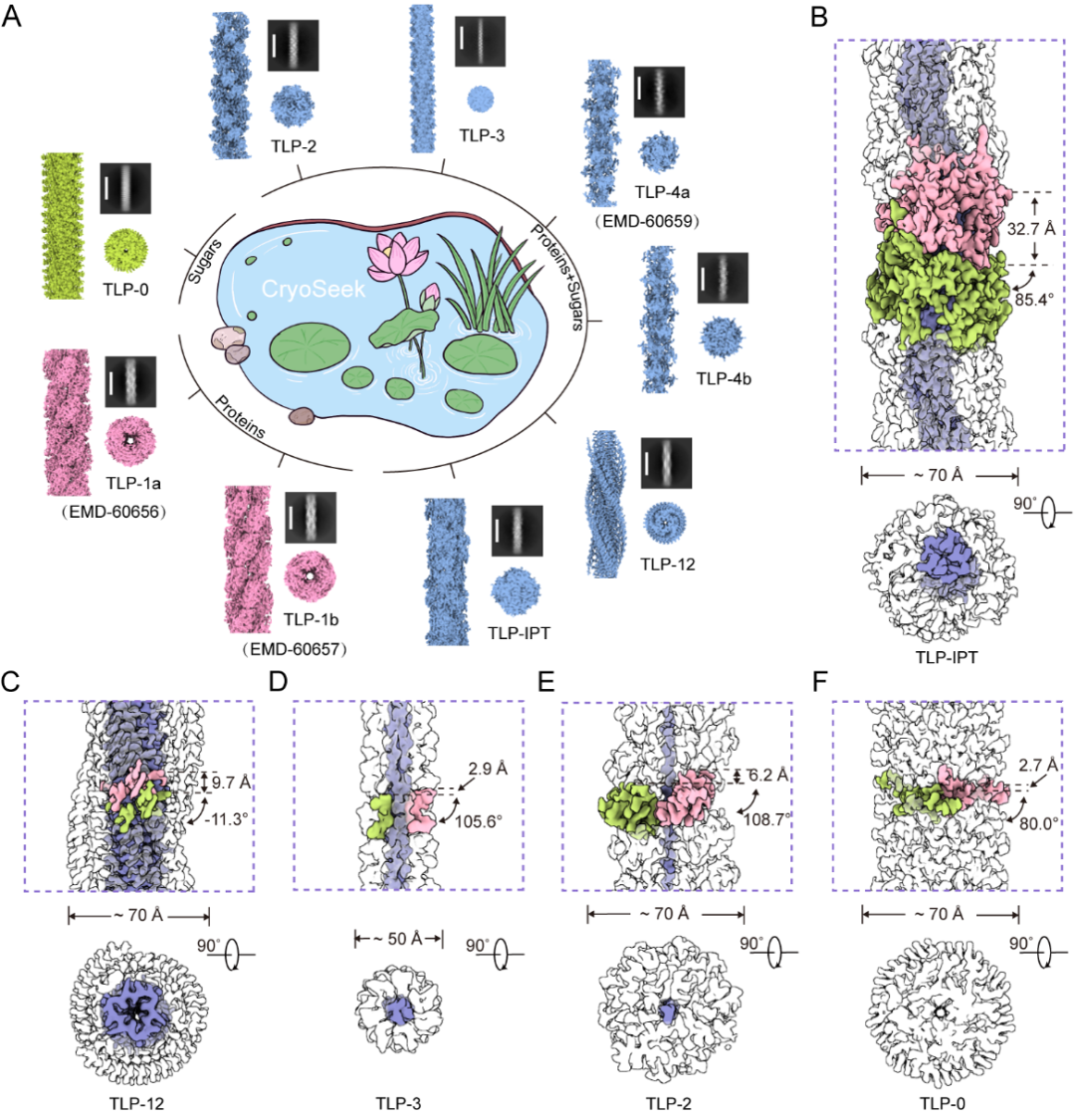
Fig. 2 CryoSeek strategy enables identification of various fibrils in water samples from Tsinghua lotus pond
Fig. 2A To date, nine fibrils have been identified at high resolution in water samples from Tsinghua lotus pond. Based on protein content, they are categorized into three types: protein fibrils (TLP-1a/b), glycofibrils with protein cores (TLP-2/3/4a/4b/12/IPT), and a protein-free glycofibril (TLP-0);
Figs. 2B-F Composition and helical parameters of the five newly identified glycofibrils
The five newly identified glycofibrils are named as TLP-IPT, TLP-12, TLP-3, TLP-2, and TLP-0. The prefix “TLP” stands for Tsinghua Lotus Pond, while the suffix denotes features of the protein core within its structure. TLP-IPT contains a recognizable protein core formed by consecutive IPT (Ig‐like/plexins/transcription factors) domains, each encircled by 13 glycan chains. TLP-12 is made of three highly repeated dodecapeptide strands that intertwine into a triple parallel β‐sheet ribbon, overlaid by two helical glycan ridges. Both TLP-3 and TLP-2 possess linear polypeptide core: TLP-3 is formed by three intertwined tripeptide repeats , whereas TLP-2 is composed of a single dipeptide repeat. Each is extensively coated by a thick layer of glycans. TLP-0 is composed entirely of glycans. The suffix “0” indicates the absence of any protein component. These findings not only reveal the structural and compositional diversity of natural glycofibrils but also demonstrate how CryoSeek strategy, integrated with multidisciplinary approaches, enables high-throughput, systematic glycan sequencing and three-dimensional structural determination.
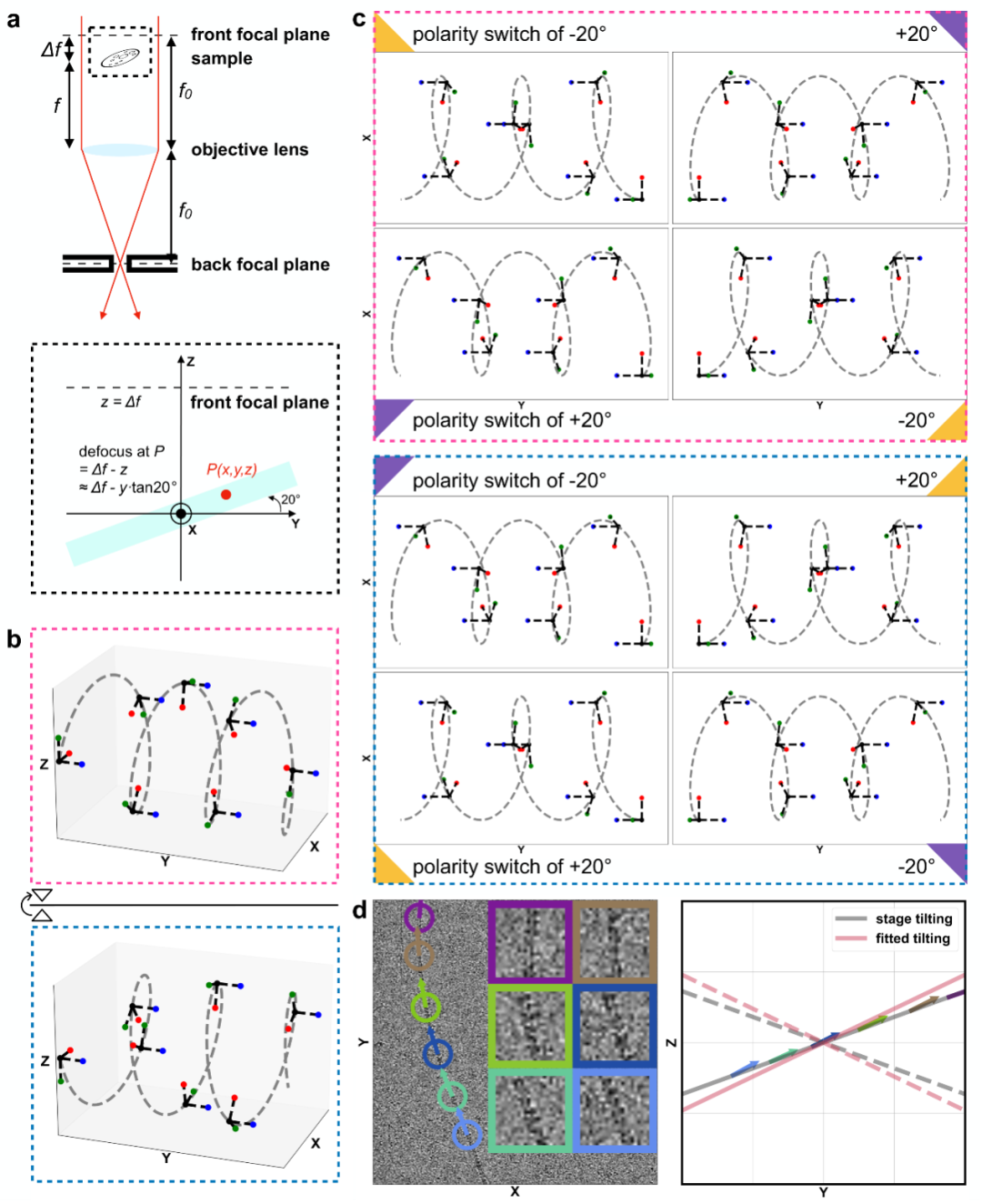
Fig. 3 Ahaha principle for absolute hand determination
To determine the absolute hand of glycofibrils, the team developed the Ahaha algorithm. As illustrated in Fig. 3, its principle rests on the fact that enantiomeric helices yield different projections at the same tilt angle, yet identical projections at opposite tilts. Thus, helices of opposite absolute hand exhibit opposite tilt preferences. Furthermore, as these glycofibrils lie flat within the sample plane, their tilt is constrained to align with it. Ahaha determines the correct absolute hand of a high-resolution reconstructed density map by assessing whether the tilt orientations of individual particles are consistent with the overall sample tilt. Applied to a natural freshwater sample (dripping stalagmite in a karst cave), Ahaha successfully determined the absolute hand of four glycofibrils and enabled the accurate reconstruction of their atomic models (Fig. 4). This represents a major milestone in determining the structure of the glycan, one "dark matter" of life. The launch of this online service (https://cryoseek.org/ahaha) offers a powerful tool to researchers worldwide and holds significant potential for accelerating breakthroughs in glycobiology.
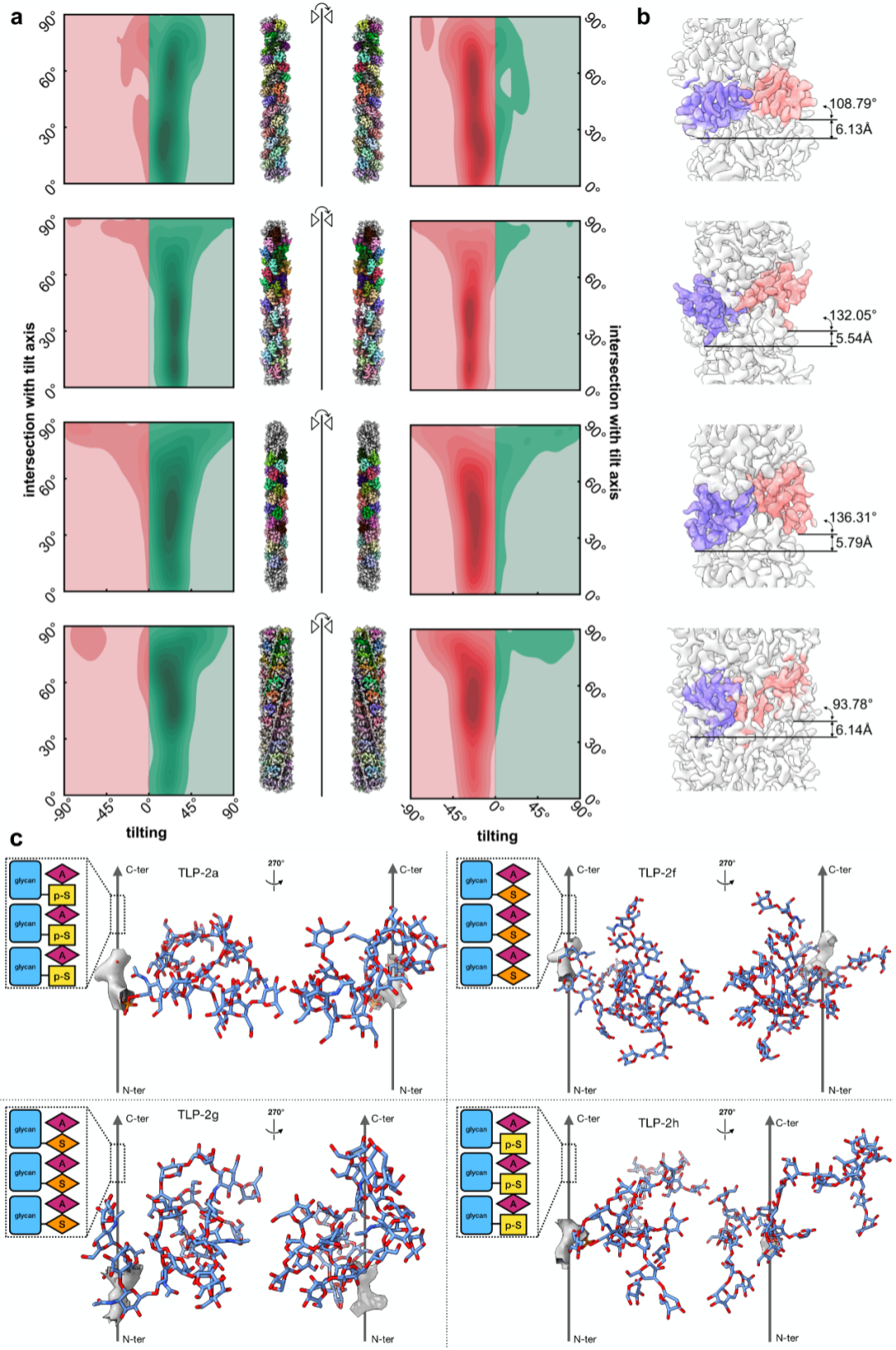
Fig. 4 Ahaha determines the absolute hand and enables atomic modeling of four natural glycofibrils
Fig. 4a Heatmaps from Ahaha analysis of particle tilt orientations. Alignment with the sample tilt (green) indicates correct absolute hand; misalignment (red) indicates the incorrect absolute hand. Refinements using initial models with inverted helical twist and absolute hand yield incorrect results as anticipated
Fig. 4b Two adjacent helical units of glycofibrils and their helical symmetry parameters
Fig. 4c Atomic models of glycofibrils
The first study was led by co-corresponding authors Nieng Yan (Professor, Shenzhen Medical Academy of Research and Translation & Shenzhen Bay Laboratory), Zhangqiang Li (Research Assistant, School of Life Sciences, Tsinghua University), and Tongtong Wang (PhD, School of Life Sciences, Tsinghua University). Zhangqiang Li (Research Assistant), Tongtong Wang (PhD), and Yitong Sun from the School of Life Sciences at Tsinghua University are the co-first authors. Other contributors include Kui Xu (PhD), Wenze Huang (PhD), Qiangfeng Zhang (Associate Professor), and Chuangye Yan (Associate Professor) from School of Life Sciences at Tsinghua University, as well as Mingxu Hu (Junior Principal Investigator, Shenzhen Medical Academy of Research and Translation). The second study was led by co-corresponding authors Mingxu Hu (Junior Principal Investigator, Shenzhen Medical Academy of Research and Translation), Jiawei Wang (Associate Professor, School of Life Sciences, Tsinghua University), and Nieng Yan (Professor, Shenzhen Medical Academy of Research and Translation & Shenzhen Bay Laboratory). Qi Zhang (PhD, Shenzhen Medical Academy of Research and Translation) is the first author. Other contributors include Lanju Qin, Tongtong Wang (PhD), Zhangqiang Li (Research Assistant), Yilin Zhang, and Sheng Chen (PhD). The studies were funded by the Shenzhen Medical Academy of Research and Translation, National Natural Science Foundation of China, and Beijing Frontier Research Center for Biological Structure.
References:
[1] Wang, T., Li, Z., Xu, K., Huang, W., Huang, G., Zhang, Q. C., & Yan, N. (2024). CryoSeek: A strategy for bioentity discovery using cryoelectron microscopy. Proceedings of the National Academy of Sciences, 121(42), e2417046121.
[2] Wang, T., Huang, W., Xu, K., Sun, Y., Zhang, Q.C., Yan, C., Li, Z., & Yan, N. (2025). CryoSeek II: Cryo-EM analysis of glycofibrils from freshwater reveals well-structured glycans coating linear tetrapeptide repeats, Proceedings of the National Academy of Sciences, 122(1), e2423943122.
[3] Wang, T., Sun, Y., Li, Z., & Yan, N. (2024). The 8-nm spaghetti: well-structured glycans coating linear tetrapeptide repeats discovered from freshwater with CryoSeek. bioRxiv, 2024-12.